What's really inside all those nooks and crannies?
|
Many reef-associated organisms are cryptic, rare, small, or belong to poorly identified groups, making documentation of biodiversity quite challenging. With the rise of new DNA sequencing technologies and bioinformatics, community assemblages can be characterized using metabarcoding approaches from autonomous reef monitoring structures (ARMS). ARMS settlement plates were deployed across the Pacific Ocean and provide surfaces and spaces for cryptic reef organisms to settle on or shelter within. Deconstruction of ARMS units two years after deployment allow for a snapshot of biodiversity without otherwise damaging fragile coral reefs to assess cryptofauna diversity. The extraction of DNA from homogenized ARMS samples (metabarcoding) is being compared to environmental DNA (eDNA) from within and around coral reef crevices to determine community composition of micro-invertebrates that inhabit the nooks and crannies of otherwise inaccessible portions of coral reefs.
|
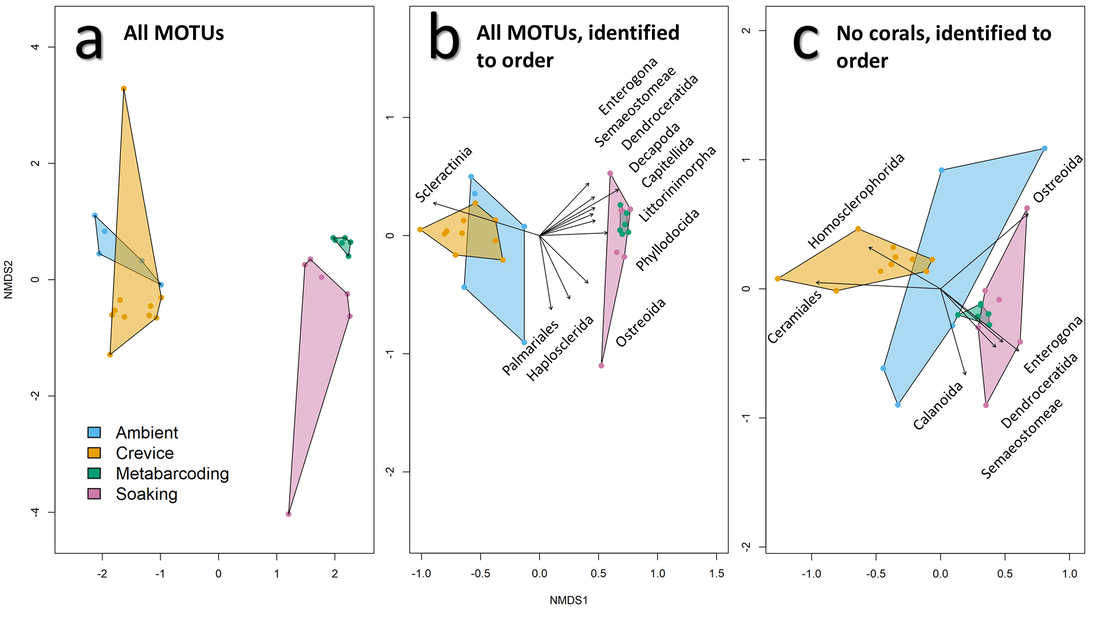
A. Samples taken from metabarcoding of ARMS settlement plates (metabarcoding) recover fundamentally different communities of cryptic coral reef taxa when compared to eDNA samples collected from the same ARMS units (soaking), the water column (ambient), and nearby reef structure (crevices). B. Ambient and crevice eDNA samples were driven largely by abundant hard corals, whereas ARMS biomass was driven largely by sponges, nematode worms, and other chitinous taxa. C. Removing corals from the analysis reveals that crevice communities were driven by sponges and red algae (Nichols et al. 2021).


